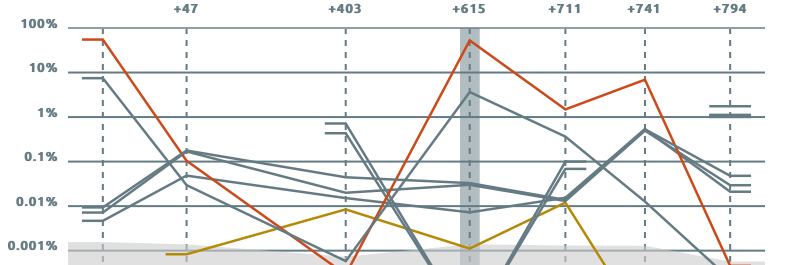
High-Throughput Analysis of V(D)J Immune Repertoire
Vidjil is an open-source platform for the analysis of high-throughput sequencing data from lymphocytes. V(D)J recombinations in lymphocytes are essential for immunological diversity. They are also useful markers of pathologies, and in leukemia, are used to quantify the minimal residual disease during patient follow-up. High-throughput sequencing (NGS/HTS) now enables the deep sequencing of a lymphoid population with dedicated Rep-Seq methods and software.
Vidjil-algo is the high-throughput algorithm of the Vidjil platform extracting V(D)J junctions and gather them into clones. This analysis is based on a seed heuristics and is fast and scalable, as, in the first phase, no alignment is performed with database germline sequences. Vidjil also contains a web application linked to a patient database for visualization and analysis of clones and their tracking along the time in a MRD setup or in a immunological study.
Availibility
Vidjil-algo is open-source, released under GNU GPLv3 license. You are welcome to redistribute it under certain conditions. This software is for research use only and comes with no warranty.
The web application can be accessed at app.vidjil.org. The development code of Vidjil-algo is available on our gitlab server, and the Vidjil-algo can also be downloaded below.
Stable releases of vidjil-algo
The full sources are in the .tgz file. The installation is documented in the doc/algo.org file.
- vidjil-algo-2018.06.tar.gz (2018/06/14), binary x86_64
- vidjil-algo-2018.02.tar.gz (2018/02/02), binary x86_64
- vidjil-2017.09.tgz (2017/09/08)
- vidjil-2017.03.tgz (2017/03/14), binary x86_64
- vidjil-2016.09.tgz (2016/09/30), binary x86_64
- vidjil-2016.08.tgz (2016/08/06)
- vidjil-2016.07.tgz (2016/07/13)
- vidjil-2016.03.tgz (2016/03/04), binary x86_64
- vidjil-2016.02.tgz (2016/02/08), binary x86_64
- vidjil-2015.12.tgz (2015/12/22), binary x86_64
- vidjil-2015.10.2.tgz (2015/10/08), binary x86_64
- vidjil-2015.07.tgz (2015/07/21), binary x86_64
- vidjil-2015.06.tgz (2015/06/05)
- vidjil-2015.05.tgz (2015/05/08)
- vidjil-2015.04.tgz (2015/04/09)
- vidjil-2015.03.tgz (2015/03/03)
- vidjil-2015.01.tgz (2015/01/31)
- vidjil-2014.12.tgz (2014/12/22)
- vidjil-2014.11.tgz (2014/11/28)
- vidjil-2014.10.tgz (2014/10/22)
- vidjil-2014.09.tgz (2014/09/23)
- vidjil-2014.07.tgz (2014/07/28)
- vidjil-2014.03.tgz (2014/03/27)
- vidjil-2014.02.tgz (2014/02/20)
- vidjil-2013.10.tgz (2013/10/07)
- vidjil-2013.07.tgz (2013/07/03)
- vidjil-2013.04.tgz (2013/04/26)
- Changelog
Documentation
- User documentation of the web application
- The documentation of the command-line version is in the doc/algo.org file that is also found in the .tgz files.
Reference
Vidjil is developed by a passionate team with fast response, from the Bonsai bioinformatics team of the CRIStAL (CNRS, U. Lille) and Inria Lille research centers, in Lille, France, and by the VidjilNet consortium. This work is in collaboration with the department of Hematology of CHRU Lille, the Functional and Structural Genomic Platform (U. Lille 2, IFR-114, IRCL), and the EuroClonality-NGS working group, and is supported by SIRIC ONCOLille (Grant INCa-DGOS-Inserm 6041), Région Nord-Pas-de-Calais (ABILES), Inria, and by Université Lille 1 (PPF Bioinformatique). The methods were presented at the JOBIM 2013 and ASH 2014 conferences, and are described in the following papers:
- Marc Duez, Mathieu Giraud, Ryan Herbert, Tatiana Rocher, Mikaël Salson, Florian Thonier, Vidjil: A web platform for analysis of high-throughput repertoire sequencing, PLOS ONE, 11(11):e0166126 (2016), doi:10.1371/journal.pone.0166126
- Yann Ferret, Aurélie Caillault, Shéhérazade Sebda, Marc Duez, Nathalie Gradel, Nicolas Duployez, Céline Villenet, Martin Figeac, Claude Preudhomme, Mikaël Salson, Mathieu Giraud, Multi-loci diagnosis of acute lymphoblastic leukaemia with high-throughput sequencing and bioinformatics analysis , British Journal of Haematology, 173(3):413–420 (2016), doi:10.1111/bjh.13981
- Mathieu Giraud, Mikaël Salson, Marc Duez, Céline Villenet, Sabine Quief, Aurélie Caillault, Nathalie Grardel, Christophe Roumier, Claude Preudhomme, Martin Figeac, Fast multiclonal clusterization of V(D)J recombinations from high-throughput sequencing, BMC Genomics, 15:409, doi:10.1186/1471-2164-15-409